The human heart beats about 115,000 times a day, pumping around 2,000 gallons of blood.
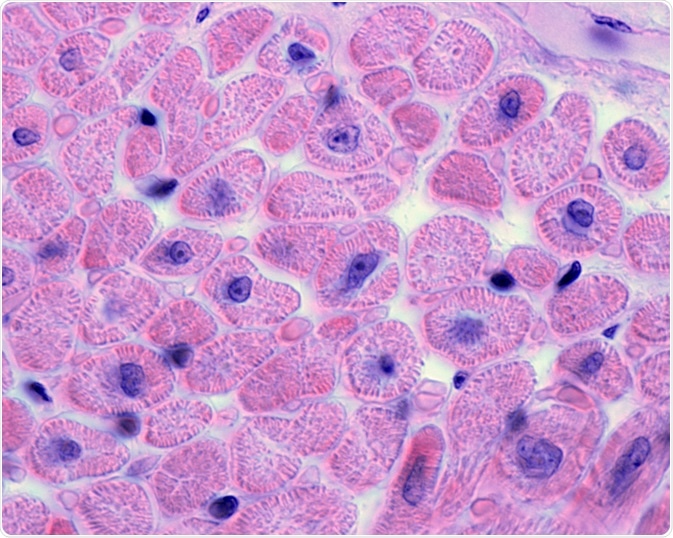
Image Credit: Jose Luis Calvo/Shutterstock.com
To enable this complex organ to perform its job effectively, without error, several specialized cell types are drawn into action.
Until recently, the diversity of cardiac cells had remained unclear. However, recent developments in scientific techniques have opened the door to gaining a more in-depth, high-resolution view of the specialized cells that make up the human heart. The differences between fetal and elderly hearts have also been highlighted lately.
Specifically, recent advances in low input RNA- sequencing technology have led to tissues being viewed clearly at the single-cell level.
Studies that demonstrated low input RNA- sequencing’s ability to define cells at such a high resolution inspired the work of a US-based team that sought to uncover the cellular diversity of cardiac tissue using this method.
A group of scientists from The Broad Institute, Massachusetts General Hospital, the University of Pennsylvania, Bayer, and the University of Groningen in the Netherlands was able to observe and define the discrete cell subtypes of the human heart using large-scale single nuclei RNA sequencing.
What the team achieved will be influential in deepening our knowledge of cardiac homeostasis and the routes to the pathology of the heart. It will likely inspire the development of next-generation therapies for cardiovascular diseases.
Using low input RNA- sequencing to define cellular diversity
The team completed single nuclear RNA sequencing in cardiac tissue samples from seven human donors using microfluidic encapsulation and barcoding. The tissue donations were from people with healthy hearts, free from cardiac disease.
Next, clusters of the individual nuclear transcriptomes were created based on the transcriptional profiles of genes with high variability.
Differential gene expression analyses, as well as analyses of the intersection with genetic and pharmacologic data of the established clusters, were run.
Data revealed 287,269 single cardiac nuclei, the transcriptomes of which were all sequenced, uncovering 9 distinct types of cells, and 20 sub-clusters of cardiac heart cells.
These sub-clusters encompassed four classes of endothelial cells, two groups of macrophages, and two groups of fibroblasts.
The comparisons of cellular transcriptomes demonstrated that diversity was evident in both cardiomyocyte transcriptional programs, as well as within the subtypes that were implicated in remodeling and vascularization processes in the extracellular matrix.
A strong enrichment was revealed by investigating the genetic association data for the role of cell subtypes in cardiac traits and diseases.
Lastly, the dataset’s intersection with genes on cardiac testing panels and the druggable genome uncovered completely new patterns of cell specificity.
Future therapies for cardiovascular disease
The research uncovered a map of the transcriptional landscape in the human heart. The conclusions reached will prove essential to developing our knowledge of how the human heart is structured.
To date, it is the largest collection of cardiac single nuclear transcriptomes allowing researchers to determine 9 major clusters and 20 sub-clusters of cell types that exist within the healthy human heart.
Also, differences between laterality-, sex-, and chamber-specific transcriptional programs were elucidated across the subtypes of cells found. Also, specific cell types were highlighted as being possibly involved in pathology due to their association with rare genetic variants underlying certain cardiovascular diseases.
Finally, the team was able to establish a method of handling cardiac single nuclear data that is portable and scalable. Future studies will be able to use its methodology to explore more tissues and hopefully gain more insights.
This data will likely influence future developments in therapeutics for a range of cardiovascular diseases.
Now that more information has been unveiled about the cell types involved in pathology, scientists are better equipped to target them. This could open the door to more effective preventions and treatment for cardiovascular diseases.
Sources:
- Arimura, T., Bos, J., Sato, A., Kubo, T., Okamoto, H., Nishi, H., Harada, H., Koga, Y., Moulik, M., Doi, Y., Towbin, J., Ackerman, M. and Kimura, A. (2009). Cardiac Ankyrin Repeat Protein Gene (ANKRD1) Mutations in Hypertrophic Cardiomyopathy. Journal of the American College of Cardiology, 54(4), pp.334-342. https://www.ncbi.nlm.nih.gov/pubmed/19608031
- Cui, Y., Zheng, Y., Liu, X., Yan, L., Fan, X., Yong, J., Hu, Y., Dong, J., Li, Q., Wu, X., Gao, S., Li, J., Wen, L., Qiao, J. and Tang, F. (2019). Single-Cell Transcriptome Analysis Maps the Developmental Track of the Human Heart. Cell Reports, 26(7), pp.1934-1950.e5. https://www.ncbi.nlm.nih.gov/pubmed/30759401
- Tucker, N., Chaffin, M., Fleming, S., Hall, A., Parsons, V., Bedi, K., Akkad, A., Herndon, C., Arduini, A., Papangeli, I., Roselli, C., Aguet, F., Choi, S., Ardlie, K., Babadi, M., Margulies, K., Stegmann, C. and Ellinor, P. (2020). Transcriptional and Cellular Diversity of the Human Heart. https://www.biorxiv.org/content/10.1101/2020.01.06.896076v1
Further Reading
Last Updated: Jan 28, 2021